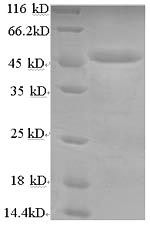
The synthesis of the recombinant plasmid containing the gene encoding the human lysosomal acid lipase/cholesteryl ester hydrolase (LIPA) protein (22-399aa) and the N-terminal GST-tag gene is the first step to produce the recombinant human LIPA protein. After that, the recombinant plasmid is transformed into E.coli cells. E.coli cells enduring a specific antibiotic are selected, demonstrating successful uptake of the recombinant plasmid. The E.coli cells containing the recombinant plasmid are cultured under conditions that encourage the expression of the gene of interest. Following expression, affinity purification is employed to isolate and purify the recombinant human LIPA protein from the cell lysate. Denaturing SDS-PAGE is applied to resolve the resulting recombinant human LIPA protein, indicating a purity level exceeding 90%.
Lysosomal acid lipase (LIPA), also known as lipoyl synthase (LipA), is a key player in several body processes. It's mainly in charge of breaking down certain fats from LDL cholesterol, helping prevent excess fat buildup [1]. LipA also helps make lipoic acid, a substance crucial for certain protein functions in the body [2][3]. It's like a builder, adding sulfur atoms to certain proteins to make them work properly [3]. Plus, it helps put together an important enzyme complex involved in energy production [4]. LipA belongs to a group of enzymes called the radical S-adenosylmethionine (SAM) superfamily, using a specific molecule to kickstart its activity [5].
Besides its role in fat metabolism, LipA is needed to make lipoic acid from scratch and might be involved in breaking down plant cell walls and sparking early immune responses [6]. In our immune cells, a gene called LAL, which makes LIPA, is really important, and scientists are trying to understand exactly how it works [7]. Certain drugs that target LIPA have shown promise in reducing fat buildup in certain diseases [8]. Also, LIPA might have a role in the formation of abnormal protein deposits seen in diseases like Parkinson's [9].
References:
[1] J. Dubland and G. Francis, Lysosomal acid lipase: at the crossroads of normal and atherogenic cholesterol metabolism, Frontiers in Cell and Developmental Biology, vol. 3, 2015. https://doi.org/10.3389/fcell.2015.00003
[2] M. Schonauer, A. Kastaniotis, V. Kursu, J. Hiltunen, & C. Dieckmann, Lipoic acid synthesis and attachment in yeast mitochondria, Journal of Biological Chemistry, vol. 284, no. 35, p. 23234-23242, 2009. https://doi.org/10.1074/jbc.m109.015594
[3] M. McLaughlin, N. Lanz, P. Goldman, K. Lee, S. Booker, & C. Drennan, Crystallographic snapshots of sulfur insertion by lipoyl synthase, Proceedings of the National Academy of Sciences, vol. 113, no. 34, p. 9446-9450, 2016. https://doi.org/10.1073/pnas.1602486113
[4] J. Miller, R. Busby, S. Jordan, J. Cheek, T. Henshaw, G. Ashleyet al., escherichia coli lipa is a lipoyl synthase: in vitro biosynthesis of lipoylated pyruvate dehydrogenase complex from octanoyl-acyl carrier protein, Biochemistry, vol. 39, no. 49, p. 15166-15178, 2000. https://doi.org/10.1021/bi002060n
[5] E. McCarthy and S. Booker, Destruction and reformation of an iron-sulfur cluster during catalysis by lipoyl synthase, Science, vol. 358, no. 6361, p. 373-377, 2017. https://doi.org/10.1126/science.aan4574
[6] A. Gudlur, A. Chatterjee, R. Sonti, & R. Sankaranarayanan, A cell wall–degrading esterase of xanthomonas oryzae requires a unique substrate recognition module for pathogenesis on rice, The Plant Cell, vol. 21, no. 6, p. 1860-1873, 2009. https://doi.org/10.1105/tpc.109.066886
[7] H. Zhang, J. Shi, M. Hachet, C. Xue, R. Bauer, H. Jianget al., Crispr/cas9-mediated gene editing in human ipsc-derived macrophage reveals lysosomal acid lipase function in human macrophages—brief report, Arteriosclerosis Thrombosis and Vascular Biology, vol. 37, no. 11, p. 2156-2160, 2017. https://doi.org/10.1161/atvbaha.117.310023
[8] K. Subramanian, N. Rauniyar, M. Lavallée-Adam, J. Yates, & W. Balch, Quantitative analysis of the proteome response to the histone deacetylase inhibitor (hdaci) vorinostat in niemann-pick type c1 disease, Molecular & Cellular Proteomics, vol. 16, no. 11, p. 1938-1957, 2017. https://doi.org/10.1074/mcp.m116.064949
[9] M. Bérard, R. Sheta, S. Malvaut, R. Rodríguez-Aller, M. Teixeira, W. Idiet al., A light-inducible protein clustering system for in vivo analysis of α-synuclein aggregation in parkinson disease, Plos Biology, vol. 20, no. 3, p. e3001578, 2022. https://doi.org/10.1371/journal.pbio.3001578"